Research Highlights
Highly efficient genome engineering in flowering plants ~ Development of a rapid method to knockout genes in Arabidopsis thaliana ~
Plant biologists at ITbM, Nagoya University have developed a genome editing method to knockout target genes in a model plant with high efficiency. The team reports a new CRISPR/Cas9 vector for the model plant that can strongly induce inheritable mutations. This method is expected to become a powerful molecular tool for genome engineering in various plant species.
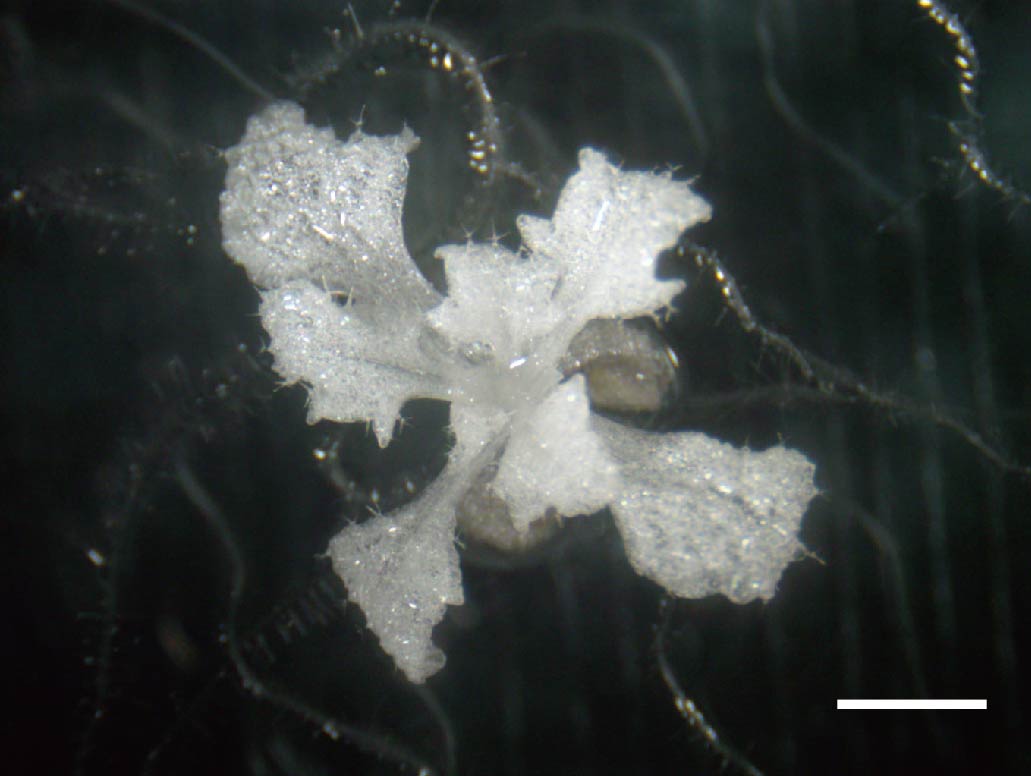

An Arabidopsis thaliana plant turned into an albino species by knockout of the PDS3 gene. Successful expression of the Cas9 protein with an RPS5A promoter (pKIR vector) led to knockout of the PDS3 gene, which is responsible for chlorophyll synthesis. Scale bar is 2 mm.
About the research:
Nagoya, Japan - A pair of plant biologists at the Institute of Transformative Bio-Molecules (ITbM) of Nagoya University, has reported in the journal Plant and Cell Physiology, on the development of a new vector (a carrier to transfer genetic information) to knockout the target genes in the model plant, Arabidopsis thaliana, in a highly efficient and inheritable manner.
The genome consists of the organism's complete set of DNA, including its genes, which contains all the information needed to develop and maintain the organism. Genome engineering, which involves specific modification of parts of the genome by removing, adding and altering sections of the DNA sequence, is a rapidly developing technique. So far, the CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats)/Cas9 (CRISPR-associated protein 9) system is one of the most popular methods for genetic manipulation arising from its simplicity, versatility and efficiency.
Nonetheless, the mutation inducing efficiency of CRISPR/Cas9 towards the model plant, Arabidopsis thaliana, has remained somewhat low so far. This is because the trigger for gene mutation is activated at later developmental stages in cells. Therefore, a significant amount of time, effort and plant species has been required to obtain the desired plant species with the targeted gene knocked out.
CRISPR/Cas9 consists of 2 key molecules, a piece of RNA called guide RNA (gRNA) and an enzyme called Cas9. Cas9 acts as a molecular scissor that can cut both strands on DNA at a specific location in the genome, so that DNA sequences can be added or removed at that position. On the other hand, the gRNA consists of a pre-designed RNA sequence (about 20 bases long) that is located within a longer RNA scaffold. The RNA scaffold binds to DNA and the pre-designed RNA sequence guides the Cas9 protein to the specific part of the genome, so that Cas9 can cut at the targeted position.
"By using a promoter for a RIBOSOMAL PROTEIN S5A (RPS5A) gene, which is expressed from an early embryonic stage in plant cells, we were able to induce the Cas9 protein to knockout genes in egg cells with high efficiency," says Hiroki Tsutsui, the first author of this study. "This RPS5A promoter is active in egg cells and we decided to call this molecular tool, a pKAMA-ITACHI Red (pKIR) vector, which can edit the plant's genome in high efficiency relative to the 35S promoter commonly used in plants," he continues.
'Kama-itachi' has the Japanese meaning of three weasel-like creatures, where the first weasel targets and trips a person, followed by the second one injuring the person by cutting them, and the third one healing the person. This is similar to the process of the CRISPR/Cas9 system towards DNA, where CRISPR targets, Cas9 cuts, and induces repair of the specific DNA sequence of interest.
"By being able to efficiently knockout the targeted gene in Arabidopsis thaliana, we consider this to be a promising method to elucidate the genetic functions of plants," says Tetsuya Higashiyama, a Professor and leader of this research. "We hope that we can apply this methodology for genome editing of crops, such as Brassica napus, to accelerate their growth and generate a variety of plant lines."
The CRISPR/Cas9 system works by knocking out a specific gene in order to investigate their function. In the model plant, Arabidopsis thaliana, as the Cas9 protein is expressed at a later developmental stage of the cell, the degree of gene knockout varies according to the tissue. The genome mutation efficiency has therefore been relatively low.
For instance, when a commonly used 35S promoter for plants was used to express the Cas9 protein in Arabidopsis thaliana, although frequent knockouts of the genes were observed in the leaves, only a few were detected in flowers. This suggests that the knockout mutation efficiency of the target gene was relatively low in the reproductive cells of flowers. Subsequently, the knockout mutation was difficult to be passed on to the daughter cells in the next generation.
In order to solve this issue, Higashiyama's group decided to express the Cas9 protein in the egg cell and in the cell during the early developmental stage, in order to improve the knockout efficiency of the genes.
"Since egg cells and fertilized egg cells (zygotes) are the origin for plant cells to develop and grow, we figured that if genome editing is carried out at an early stage, gene mutation may occur with high efficiency and can be inherited by the next generation of cells," explains Tsutsui. "The pKIR vector was ideal as it can continuously express the Cas9 protein during the egg cell stage and the developmental stage."
The team first attempted the knockout of the PDS3 (Phytoene DeSaturase 3) gene, which is known to be responsible for the synthesis of chlorophyll in plants. Without chlorophyll, the plant will become an albino species with a white appearance. By expressing Cas9 with pKIR, Tsutsui succeeded in observing the knockout of the PDS3 gene, indicated by the generation of an albino plant.
Tsutsui also investigated the effect on the amount of chlorophyll synthesized in the flower stem, when the PDS3 gene was knocked out. Upon use of the 35S promoter to express Cas9, the amount of chlorophyll was only slightly different to the amount observed in the wild type. On the other hand, when Cas9 was induced by pKIR, the amount of chlorophyll decreased drastically, suggesting that knockout of the gene had occurred efficiently.
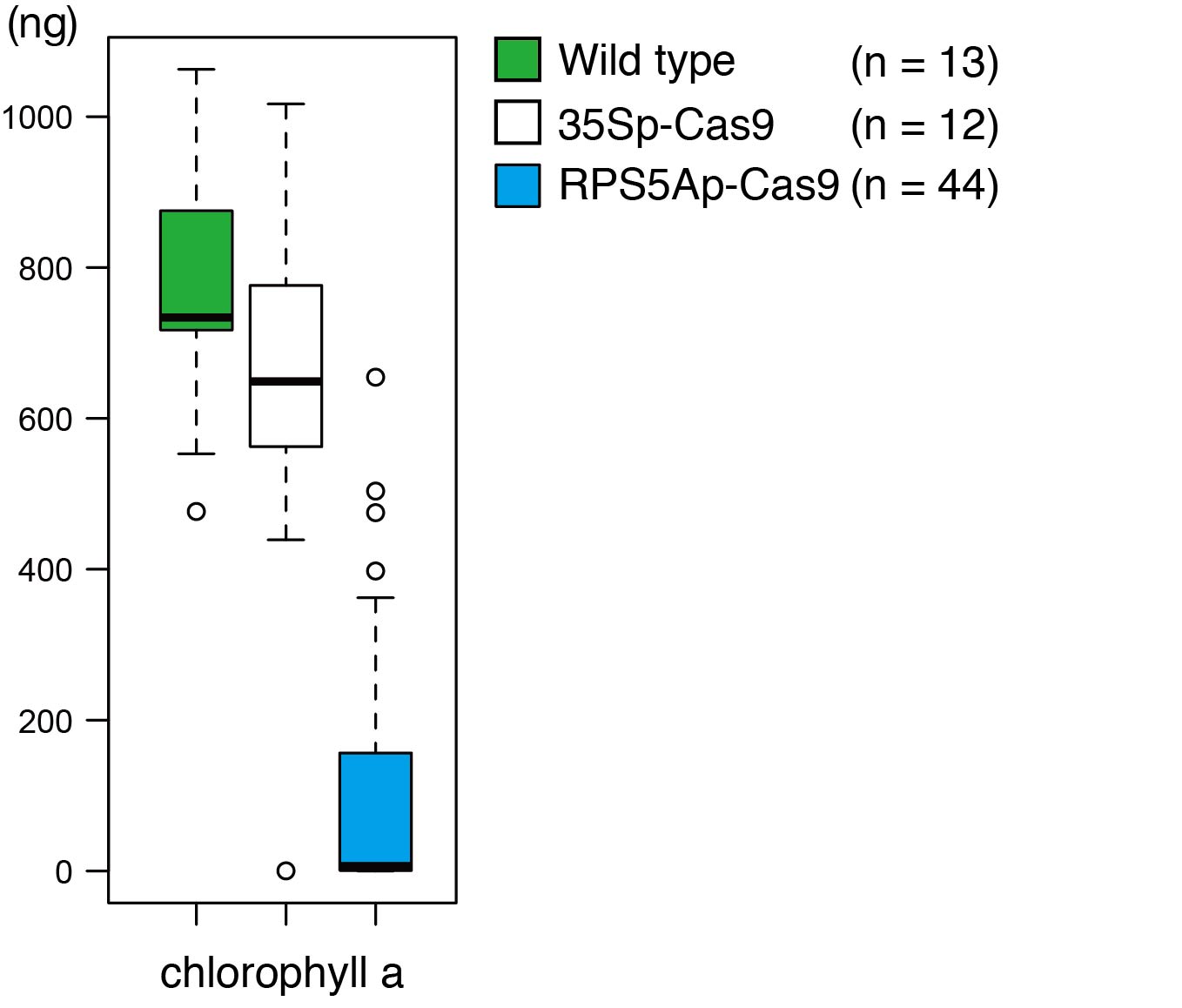
The amount of chlorophyll in the flower stems of Arabidopsis thaliana with knockout of the PDS3 gene. The amount of chlorophyll a in 1 mg of flowered stem. When the Cas9 protein was expressed by a 35S promoter (35Sp-Cas9), the amount of chlorophyll a did not differ much with the wild type. When Cas9 was expressed by the RPS5A promoter (RPS5Ap-Cas9), the amount of chlorophyll a decreased drastically. n represents the number of plant species investigated.
The team also found that the pKIR vector successfully knocks out other genes in Arabidopsis thaliana, such as AGAMOUS (AG), DUO1 (DUO POLLEN 1), and ADH1 (ALCOHOL DEHYDROGENASE 1), which are involved in the development of the flower, sperm cell, and an enzyme that converts alcohol to aldehydes, respectively.
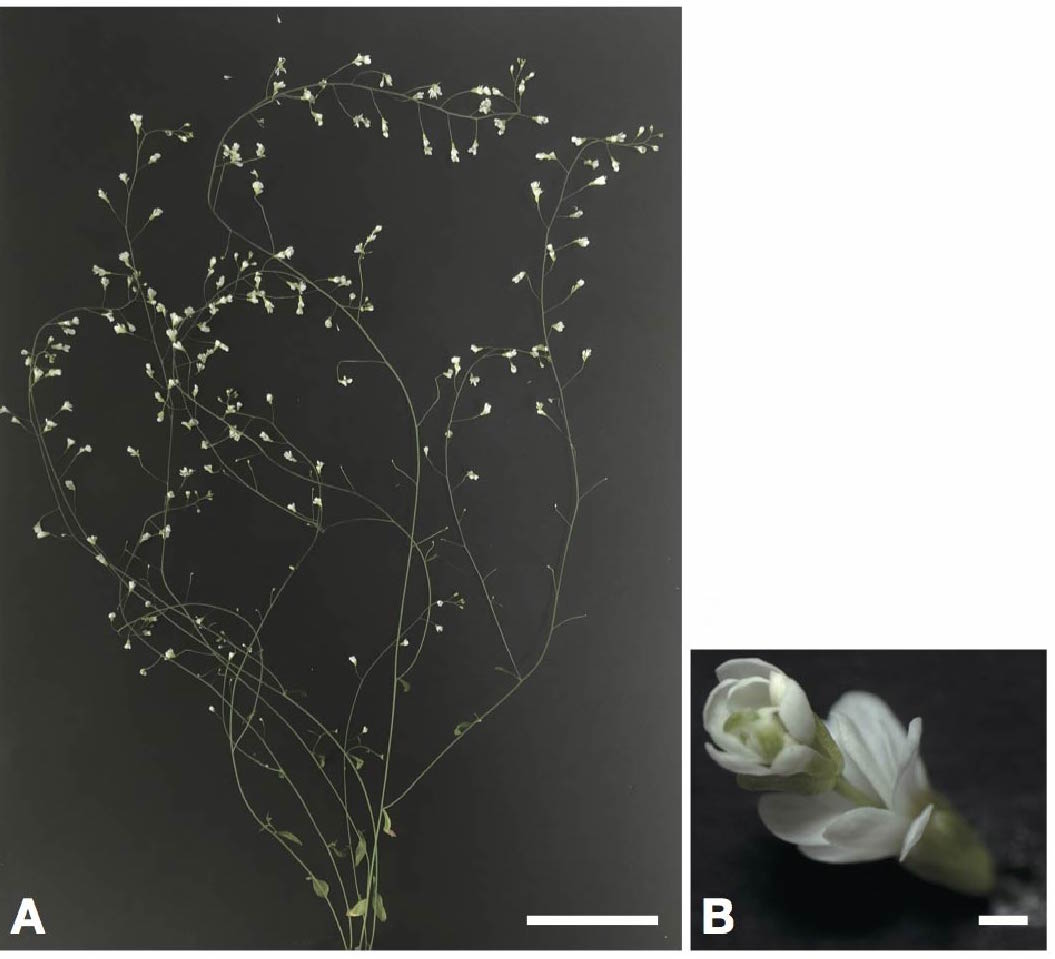

Photo of Arabidopsis thaliana with knockout of the AGAMOUS gene. (A) Observation of a flower within a flower (double flower). Scale bar is 5 cm; (B) Enlarged photo of the double flower. Scale bar is 1 cm.
AGAMOUS is a transcription factor involved in flower development (floral meristem). Upon knockout of the AGAMOUS gene induced by pKIR, a flower within a flower, known as a double flower was observed in high frequency. In addition, in the knockout experiment of DUO1 gene, which encodes a male germ line, induced by pKIR, cell division of the generative cell was impaired and a non-fertile sperm-like cell was observed.
The ADH1 gene, which encodes an enzyme that converts allyl alcohol into acrolein (an aldehyde), was also knocked out by induction with pKIR in high frequency. This high frequency suggests that the ADH1 gene was knocked out in all of the reproductive cells (egg and sperm cells) in the first generation. Mutants with the ADH1 gene knockout were able to survive, while the wild type and heterozygous plants were killed due to the generation of acrolein, which is toxic for plants.
"Since all of the species in the second generation showed the same mutation pattern, this indicates that genome editing had occurred early in the developmental stage (in the egg cells of the first generation)," describes Tsutsui.
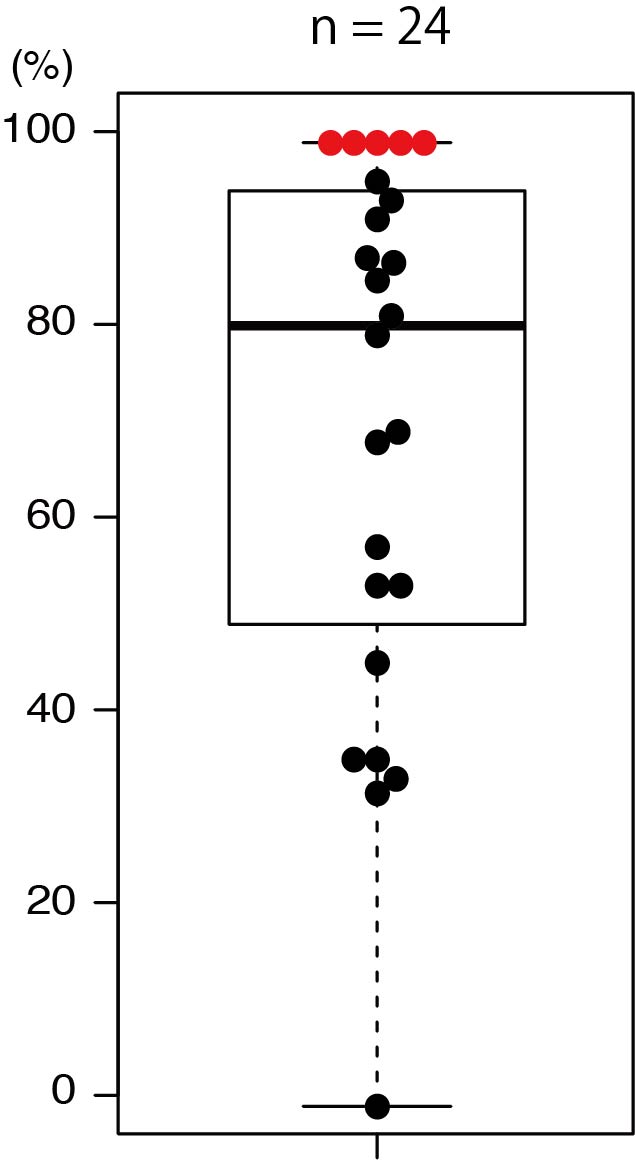
Proportion of Arabidopsis thaliana knockout of the ADH1 gene. Upon investigating the proportion of knockout in 24 lines of the 2nd generation, the median was 81% and 5 lines had 100% knockout mutation.
In order to easily identify the species installed with genes for CRISPR/Cas9, the seeds that contain Cas9 are marked by red fluorescence.
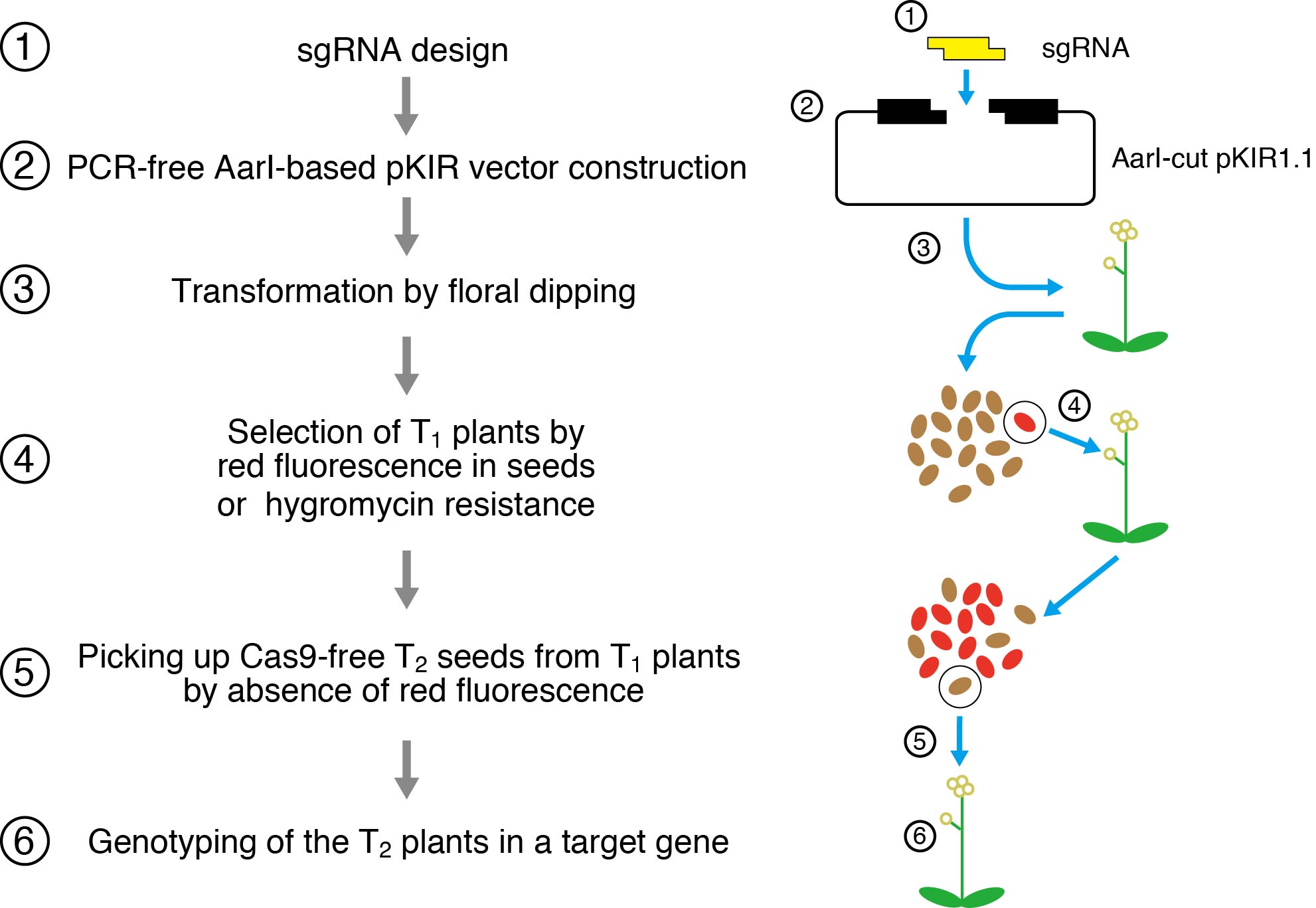
Experimental procedure for gene knockout using the pKIR vector. Plant seeds containing Cas9 are marked with a red fluorescent marker to tell which seeds contain Cas9. Using this method, it becomes possible to obtain seeds that have a knockout mutation in their target gene and do not contain Cas9, in the second generation by selection. This whole process takes about 3 months.
Tsutsui and Higashiyama have found an efficient method for genome editing of Arabidopsis thaliana, which consists of expressing Cas9 with a RPS5A promoter (pKIR vector) that can knockout the genes in cells during an early developmental stage and induce mutations that can be passed on to the daughter cells in the next generation.
"This pKIR method allows us to investigate gene clusters, which may have overlapping functions," says Tsutsui. "Previously, we had to make multiple knockout plants by crossing existing mutants to examine overlapping gene functions, which took time. We should be able to explore the functions of unidentified gene clusters by being able to rapidly access mutants by our relatively low cost method."
"We hope we can continue to improve this method to increase the mutation efficiency, so that it becomes a useful genome engineering tool for modifying the target DNA sequences in various organisms," says Higashiyama.
Journal Information:
This article "pKAMA-ITACHI vectors for highly efficient CRISPR/Cas9-mediated gene knockout in Arabidopsis thaliana" by Hiroki Tsutsui and Tetsuya Higashiyama is published online in Plant and Cell Physiology.
Links:


Prof. Tetsuya Higashiyama and Mr. Hiroki Tsutsui
Media Coverage and Related Links
- AlphaGalileo "Highly efficient genome engineering in flowering plants" (December 5, 2016)
- ResearchSEA "Highly efficient genome engineering in flowering plants" (December 5, 2016)
- EurekAlert! "Highly efficient genome engineering in flowering plants" (December 5, 2016)
- Science Daily "Highly efficient genome engineering in flowering plants" (December 5, 2016)
- Scienmag "Highly efficient genome engineering in flowering plants" (December 5, 2016)
- Bioengineer "Highly efficient genome engineering in flowering plants" (December 5, 2016)
- Breaking News "Highly efficient genome engineering in flowering plants" (December 5, 2016)
- SeedQuest "Highly efficient genome engineering in flowering plants" (December 5, 2016)
- Lab Manager "Highly efficient genome engineering in flowering plants" (December 5, 2016)
- Science Newsline "Highly efficient genome engineering in flowering plants" (December 5, 2016)
- Phys.Org "Development of a rapid method to knockout genes in Arabidopsis thaliana" (December 5, 2016)
- Global Plant Council "Development of a rapid method to knockout genes in Arabidopsis thaliana" (December 5, 2016)
- Genetic Literacy Project "New, more efficient method for using CRISPR to genetically edit flowering plants" (December 7, 2016)


